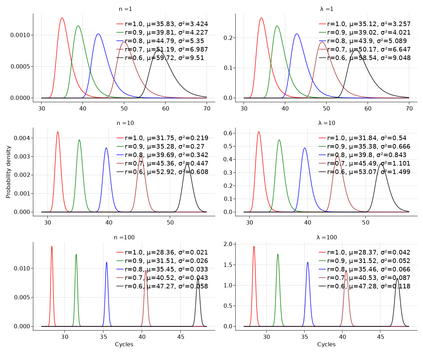
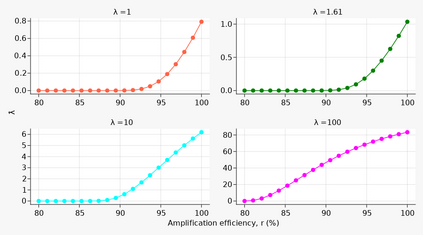
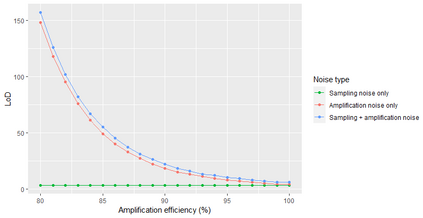
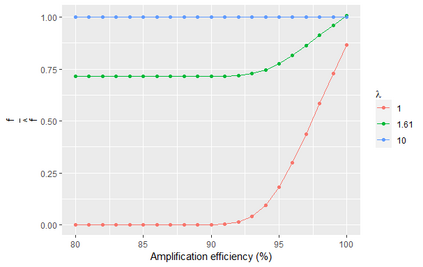
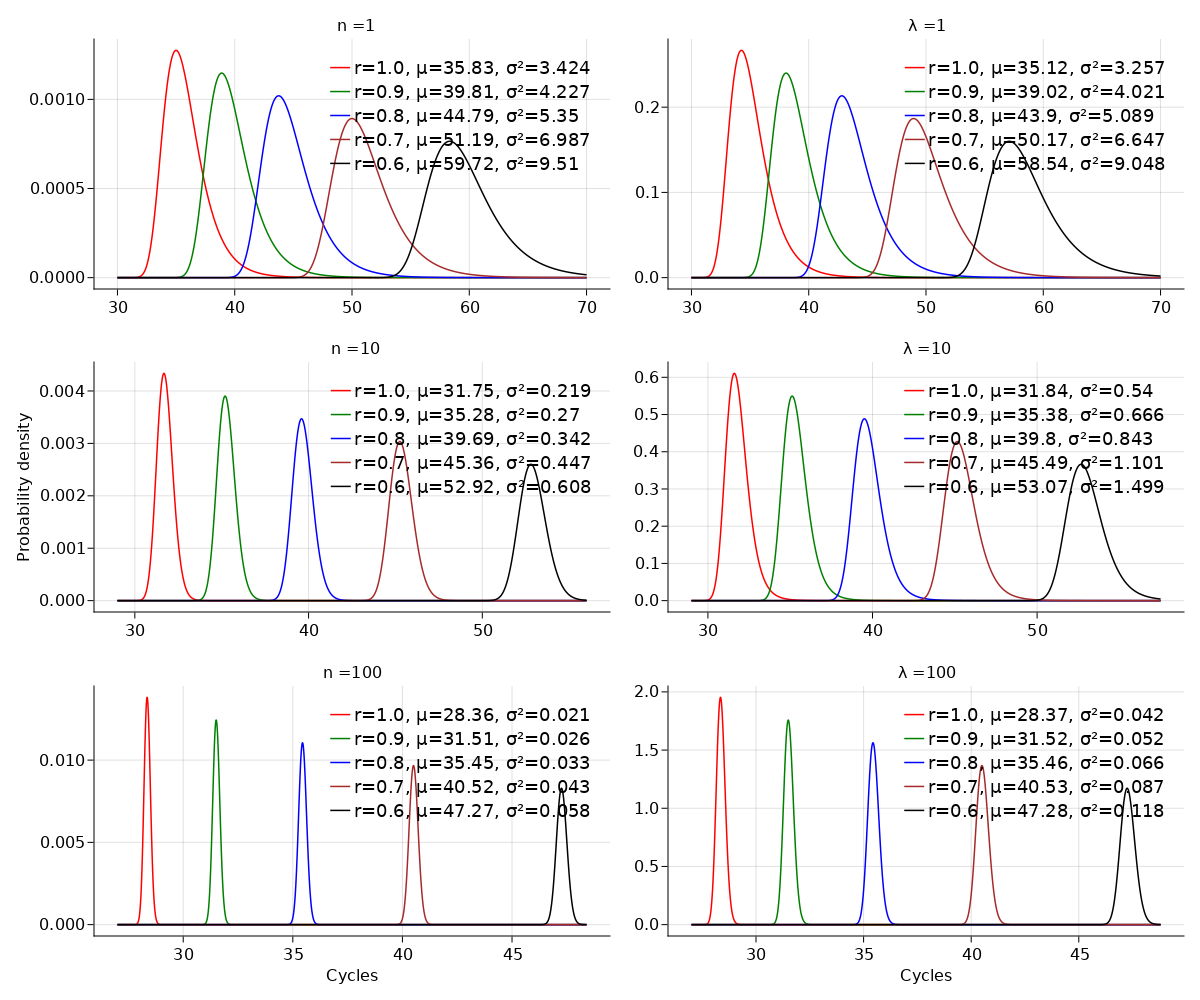
A common approach to quantifying DNA involves repeated cycles of DNA amplification. This approach, employed by the polymerase chain reaction (PCR), produces outputs that are corrupted by amplification noise, making it challenging to accurately back-calculate the amount of input DNA. Standard mathematical solutions to this back-calculation problem do not take adequate account of such noise and are error-prone. Here, we develop a parsimonious mathematical model of the stochastic mapping of input DNA onto experimental outputs that accounts, in a natural way, for amplification noise. We use the model to derive the probability density of the quantification cycle, a frequently reported experimental output, which can be fit to data to estimate input DNA. Strikingly, the model predicts that a sample with only one input DNA molecule has a $<$4% chance of testing positive, which is $>$25-fold lower than assumed by a standard method of interpreting PCR data. We provide formulae for calculating both the limit of detection and the limit of quantification, two important operating characteristics of DNA quantification methods that are frequently assessed by using ad-hoc mathematical techniques. Our results provide a mathematical foundation for the rigorous analysis of DNA quantification.
翻译:DNA量化的通用方法涉及DNA放大的反复循环。聚合酶链反应(PCR)使用的这一方法产生了因放大噪声而腐蚀的产出,使得精确反算输入DNA的数量具有挑战性。标准的计算方法没有充分考虑到这种噪音,而且容易出错。在这里,我们开发了一种对输入DNA进行分解式绘图的简单数学模型,用于实验结果,以自然的方式计算放大噪声。我们使用该模型来得出量化周期的概率密度,即经常报告的实验产出,适合数据来估计输入DNA。简而言之,该模型预测,只有一个输入DNA分子的样本的检测概率为 < 4%,比解释PCR数据的标准方法假设的值低25倍。我们提供了计算检测限度和量化限度的公式,这是经常通过应用定量技术来评估DNA定量方法的两种重要的操作特征。我们的DNA定量分析结果为DNA的精确分析提供了一种数学基础。